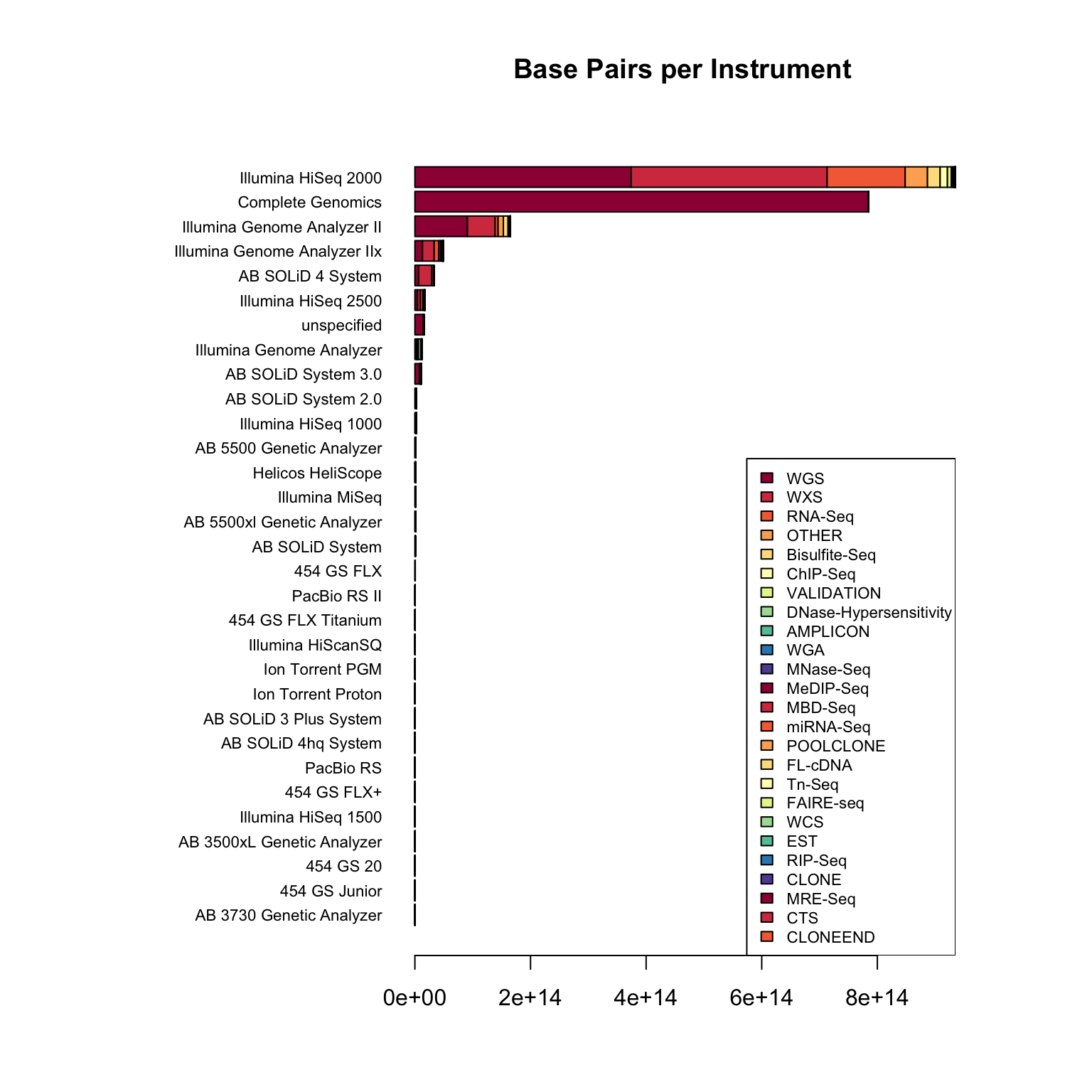Illumina Dominance in the SRA
Looking into the dominance of Illumina Sequencing in Human samples stored in the Sequence Read Archive (SRA)
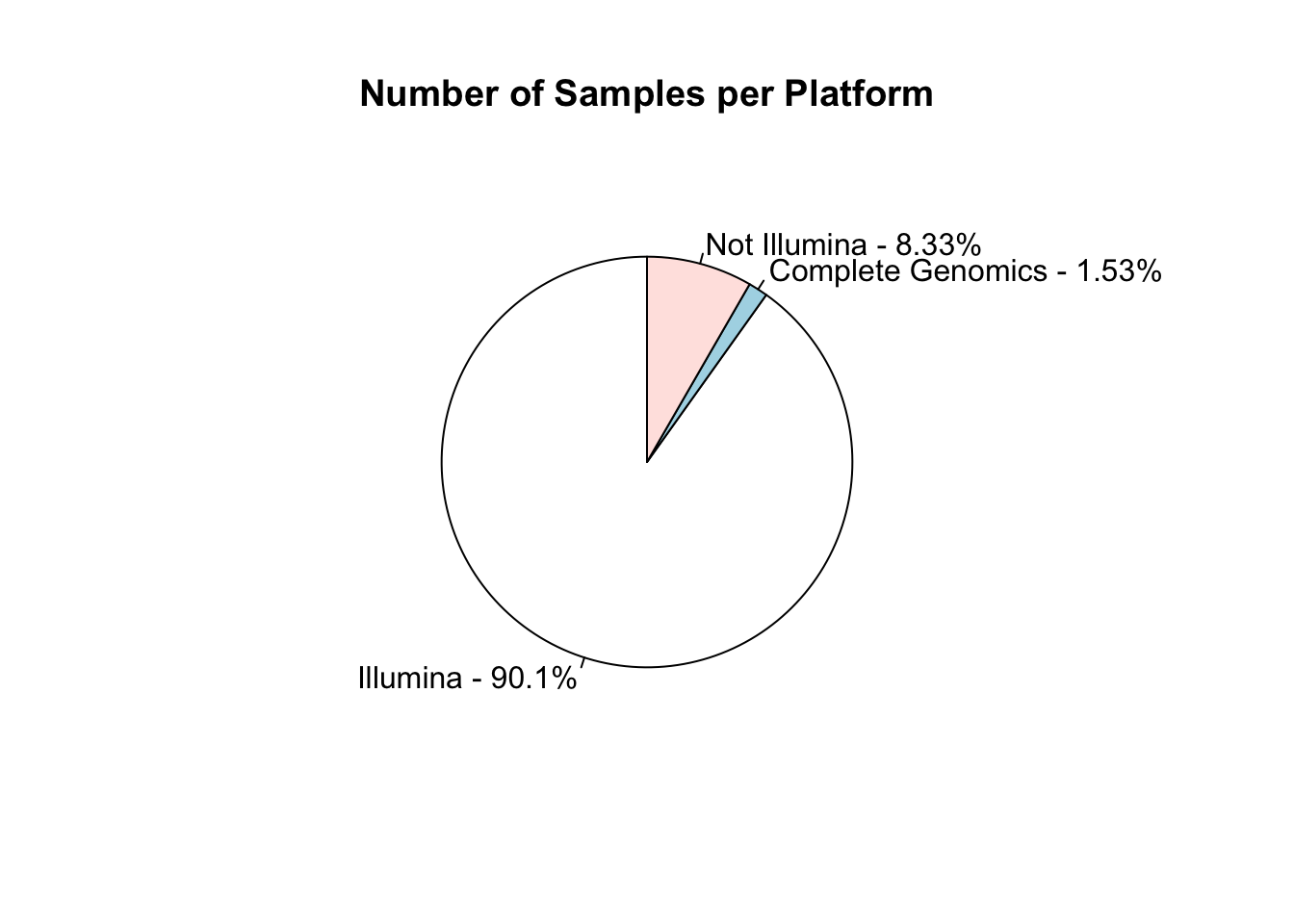
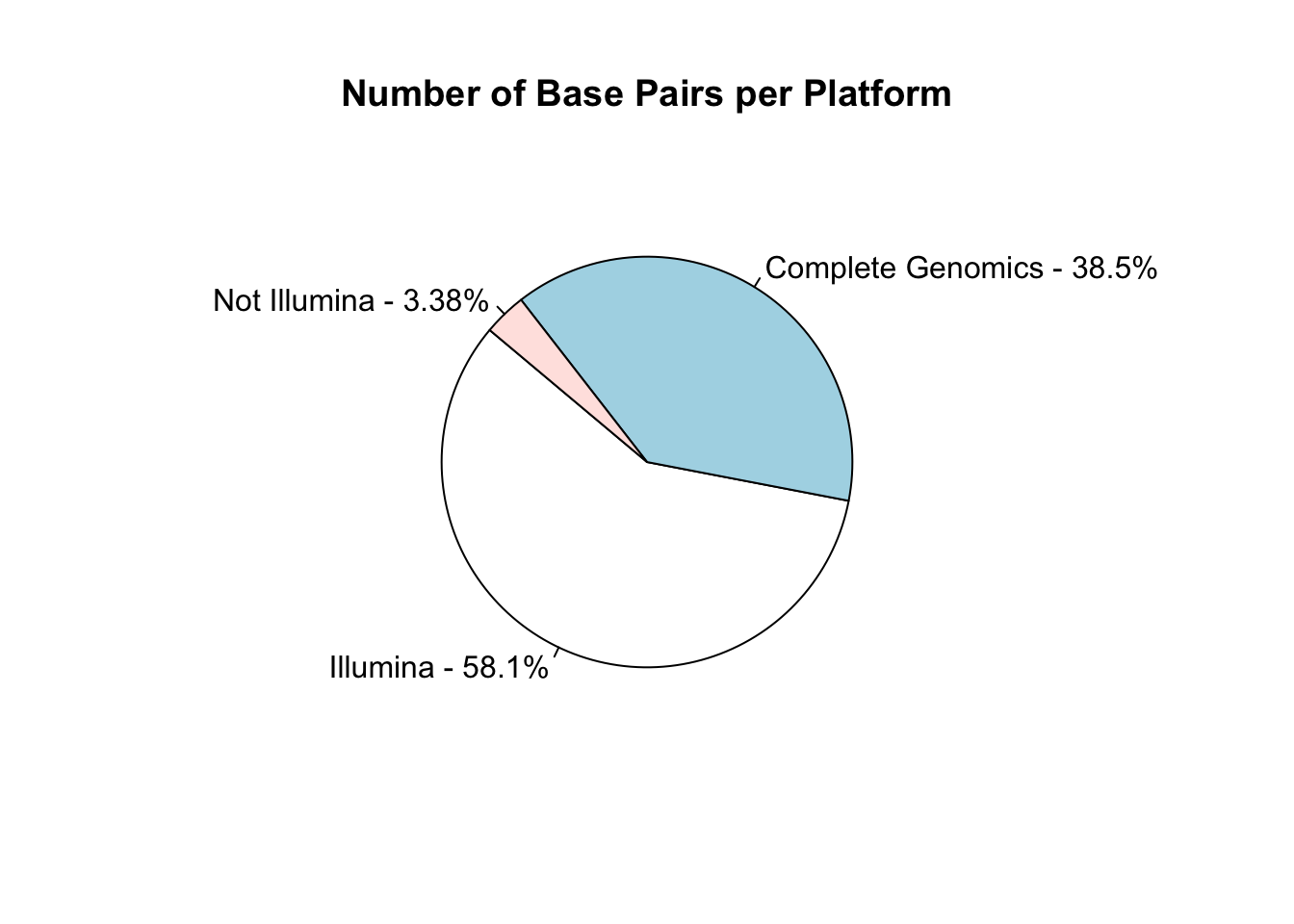
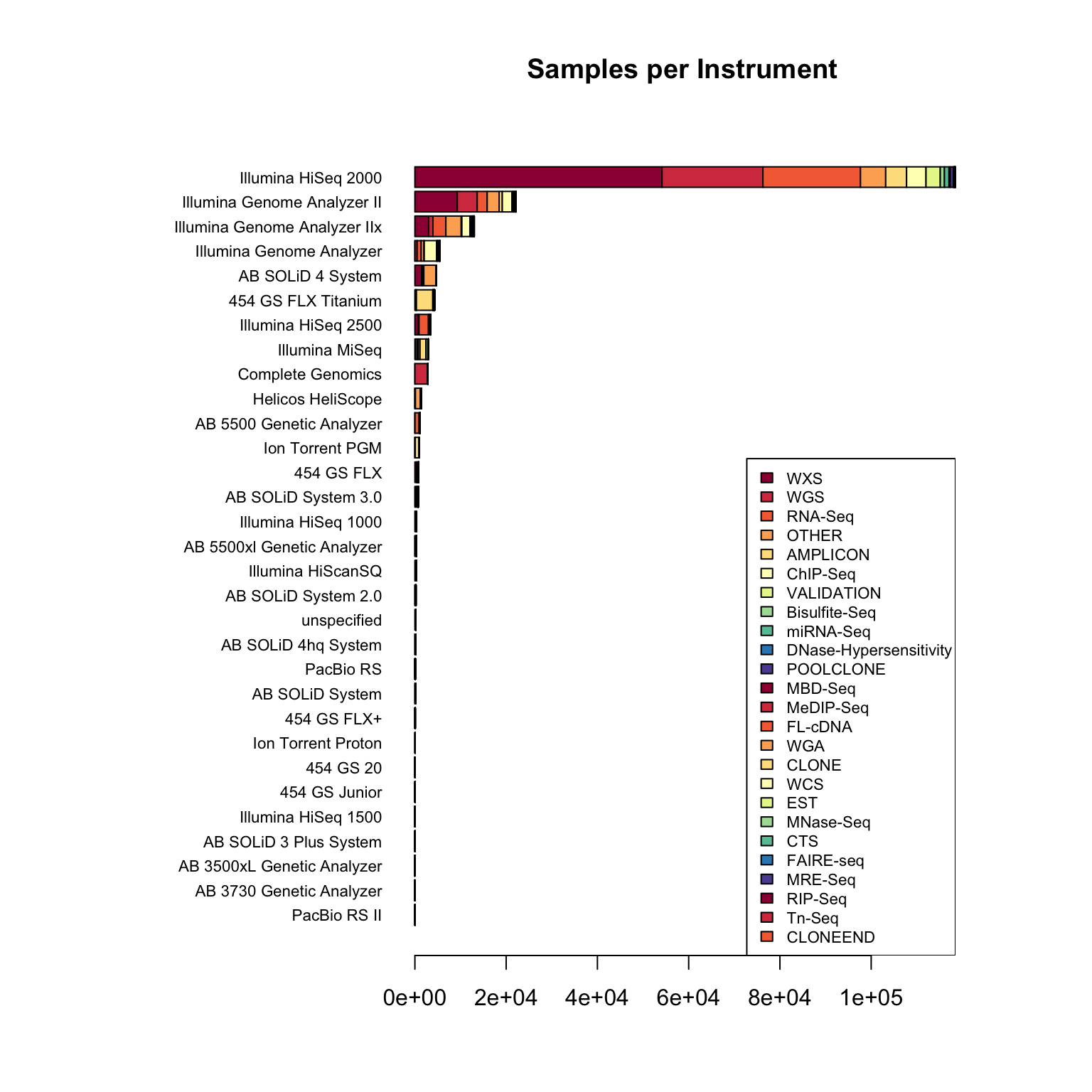
I work in biology research, specifically bioinformatics - managing data analysis for high throughput DNA sequencing. Like many labs, we use sequencing machines made by Illumina - this company is pretty dominant in the field and I was interested in just how much data has been produced on these machines.
The Sequence Read Archive is a resource where sequencing data from publications can be uploaded and made available to other researchers. I looked into the metadata associated with these samples to find out how widespread Illumina data is.
Click here to read the full knitr output from my quick R script.
Remember that this is for fun only, so don’t take it too seriously! You can see the raw script that generated it here. I’ve dropped a couple of images in below for those who want the quick overview..